SMILES Language Extension for VS Code
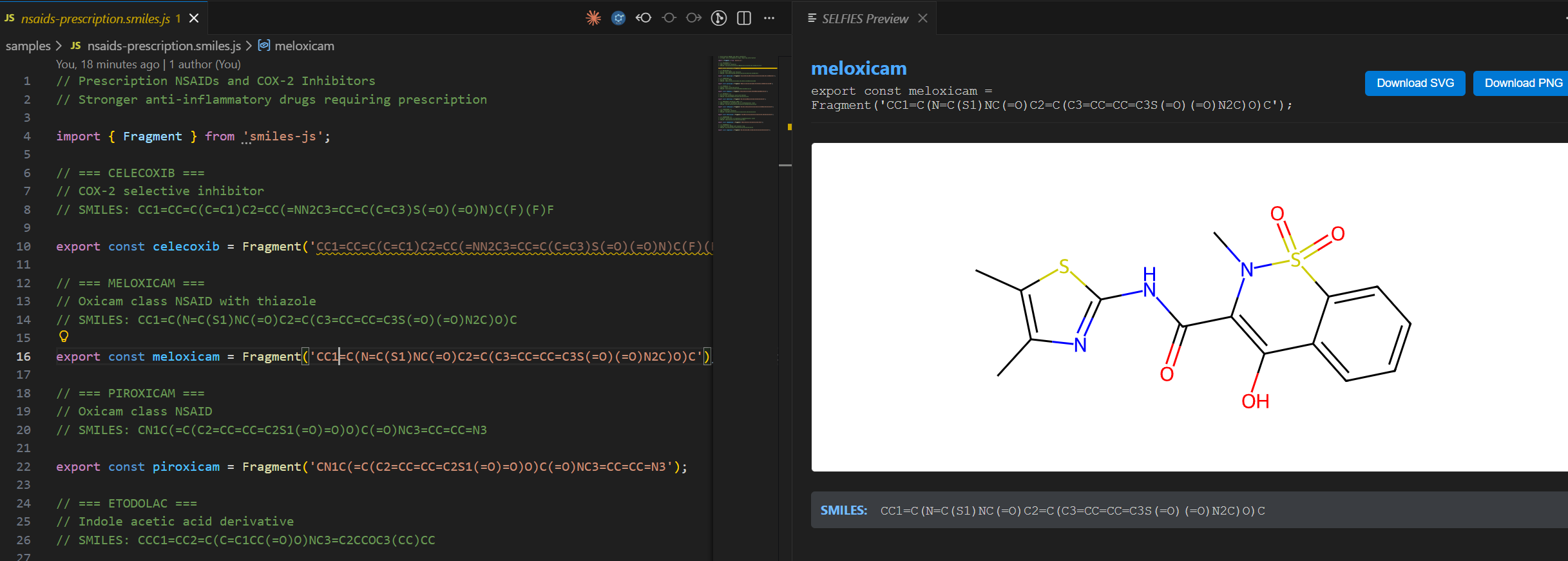
See your molecules as you write them.
Build molecules programmatically with JavaScript, compose them from reusable fragments, and watch the visualization update live as you navigate your code.
Why smiles-js?
Raw SMILES strings are write-only — CC(=O)Nc1ccc(O)cc1 tells you nothing at a glance. And when LLMs generate SMILES, they hallucinate invalid structures.
smiles-js fixes both problems: composable Fragment constructors produce valid SMILES by construction, and the code reads like chemistry:
// molecules.smiles.js
import { Fragment } from 'smiles-js';
export const methane = Fragment('C');
export const ethanol = Fragment('CCO');
export const toluene = Fragment('Cc1ccccc1');
This extension executes your JavaScript file and reads the .smiles property from exported fragments — no regex parsing! As your cursor moves, you see the 2D structure, formula, weight, and SMILES output.
Installation
Install from VS Code Marketplace
Or search "SMILES" in the VS Code extensions panel.
Quick Start
Using smiles-js (JavaScript)
- Create a file with
.smiles.js extension
- Import Fragment and Ring from
smiles-js
- Export your molecules as constants
- Move your cursor to any exported molecule to see its structure
// molecules.smiles.js
import { Fragment } from 'smiles-js';
export const methane = Fragment('C');
export const ethanol = Fragment('CCO');
export const toluene = Fragment('Cc1ccccc1');
Using SELFIES DSL
- Create a file with
.selfies extension
- Start defining molecules — the preview panel opens automatically
- Move your cursor to any definition to see its structure
# fragments.selfies
[methyl] = [C]
[ethyl] = [C][C]
[hydroxyl] = [O]
[ethanol] = [ethyl][hydroxyl]
[methanol] = [methyl][hydroxyl]
Features
smiles-js (.smiles.js files)
- JavaScript-based molecular construction
- Use Fragment() and Ring() functions
- Full JavaScript IDE support with molecular previews
SELFIES DSL (.selfies files)
- Domain-specific language for molecular composition
- Multi-file imports and reusable fragments
- Real-time diagnostics and error checking
Live Preview
As your cursor moves, the preview panel shows:
- 2D molecular structure
- Molecular formula
- Molecular weight
- SMILES output
- Export as SVG or PNG
Works for both .smiles.js and .selfies files!
Syntax Highlighting
Full highlighting for atoms, bonds, branches, rings, comments, and references in SELFIES files. JavaScript syntax highlighting for .smiles.js files.
Real-time Diagnostics
Instant feedback on SELFIES files:
- Undefined references
- Circular dependencies
- Duplicate definitions
- Syntax errors
Multi-file Projects
Use standard JavaScript imports in .smiles.js files:
import { Fragment } from 'smiles-js';
import { benzene, methyl } from 'smiles-js/common';
Or import fragments across SELFIES files:
import "./fragments.selfies" # import all
import [methyl, ethyl] from "./fragments.selfies" # import specific
Commands
SMILES: Show Molecular Structure — Open the preview panelSMILES: Toggle Preview Panel — Toggle preview on/offSMILES: Refactor Molecule to Code — Convert a Fragment to constructor code (right-click menu in .smiles.js files)
Refactor to Code (Experimental)
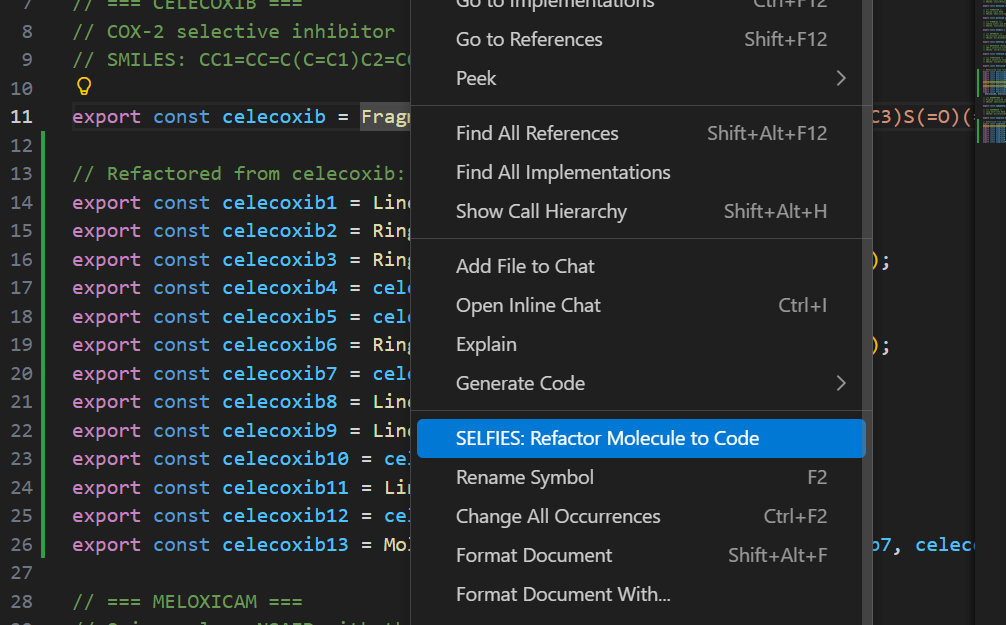
Right-click on any line with an exported Fragment in a .smiles.js file and select "SMILES: Refactor Molecule to Code" to convert it into explicit Ring/Linear constructor calls.
Note: This feature is early in development and often fails for complex molecular structures. It works best with simple fragments.
Settings
| Setting |
Default |
Description |
smiles.previewOnCursorMove |
true |
Update preview when cursor moves |
smiles.autoOpenPreview |
true |
Auto-open preview for .selfies and .smiles.js files |
smiles.renderingEngine |
smiles-drawer |
Molecule rendering engine |
Language Rules
- No forward references — Define before you use
- No recursion — No circular dependencies
- Single assignment — No redefinitions
- Case sensitive —
[Methyl] ≠ [methyl]
- smiles-js — JavaScript library for composable molecular fragments
- selfies-js — SELFIES DSL library (also usable as CLI and npm package)
- SELFIES paper — Original research by the Aspuru-Guzik group
Citation
If you use SELFIES in your research:
Krenn, M., Hase, F., Nigam, A., Friederich, P., & Aspuru-Guzik, A. (2020). Self-referencing embedded strings (SELFIES): A 100% robust molecular string representation. Machine Learning: Science and Technology, 1(4), 045024.
License
MIT



